library(DBI)
library(RMariaDB)
# Connecting to MySQL
mydb = DBI::dbConnect(RMariaDB::MariaDB(), user='msr14', password='msr14', dbname='msr14', host='localhost')
# Retrieving data from MySQL
sql <- "select user_id, follower_id from followers limit 100;"
rs = dbSendQuery(mydb, sql)
data <- fetch(rs, n=-1)26 Social Network Analysis in SE
In this example, we will data from the MSR14 challenge. Further information and datasets: http://openscience.us/repo/msr/msr14.html
Similar databases can be obtained using MetricsGrimoire or other tools.
In this simple example, we create a network form the users and following extracted from GitHub and stored in a MariaDB/MySQL database.
We can read a file directly from MySQL dump
Alternatively, we can create e CSV file directly from MySQL and load it
$mysql -u msr14 -pmsr14 msr14
> SELECT 'user','follower'
UNION ALL
SELECT user_id,follower_id
FROM followers
LIMIT 1000
INTO OUTFILE "/tmp/followers.csv"
FIELDS TERMINATED BY ','
LINES TERMINATED BY '\n';# Data already extracted and stored as CSV file (for demo purposes)
dat = read.csv("./datasets/sna/followers.csv", header = FALSE, sep = ",")
dat <- head(dat,100)We can now create the graph
library(igraph)
Attaching package: 'igraph'The following objects are masked from 'package:stats':
decompose, spectrumThe following object is masked from 'package:base':
union# Create a graph
g <- graph.data.frame(dat, directed = TRUE)Warning: `graph.data.frame()` was deprecated in igraph 2.0.0.
ℹ Please use `graph_from_data_frame()` instead.Some values:
summary(g); IGRAPH 4a5e888 DN-- 95 100 --
+ attr: name (v/c)Plotting the graph:
layout1 <- layout.fruchterman.reingold(g)
plot(g, layout1)Other layout
plot(g, layout=layout.kamada.kawai)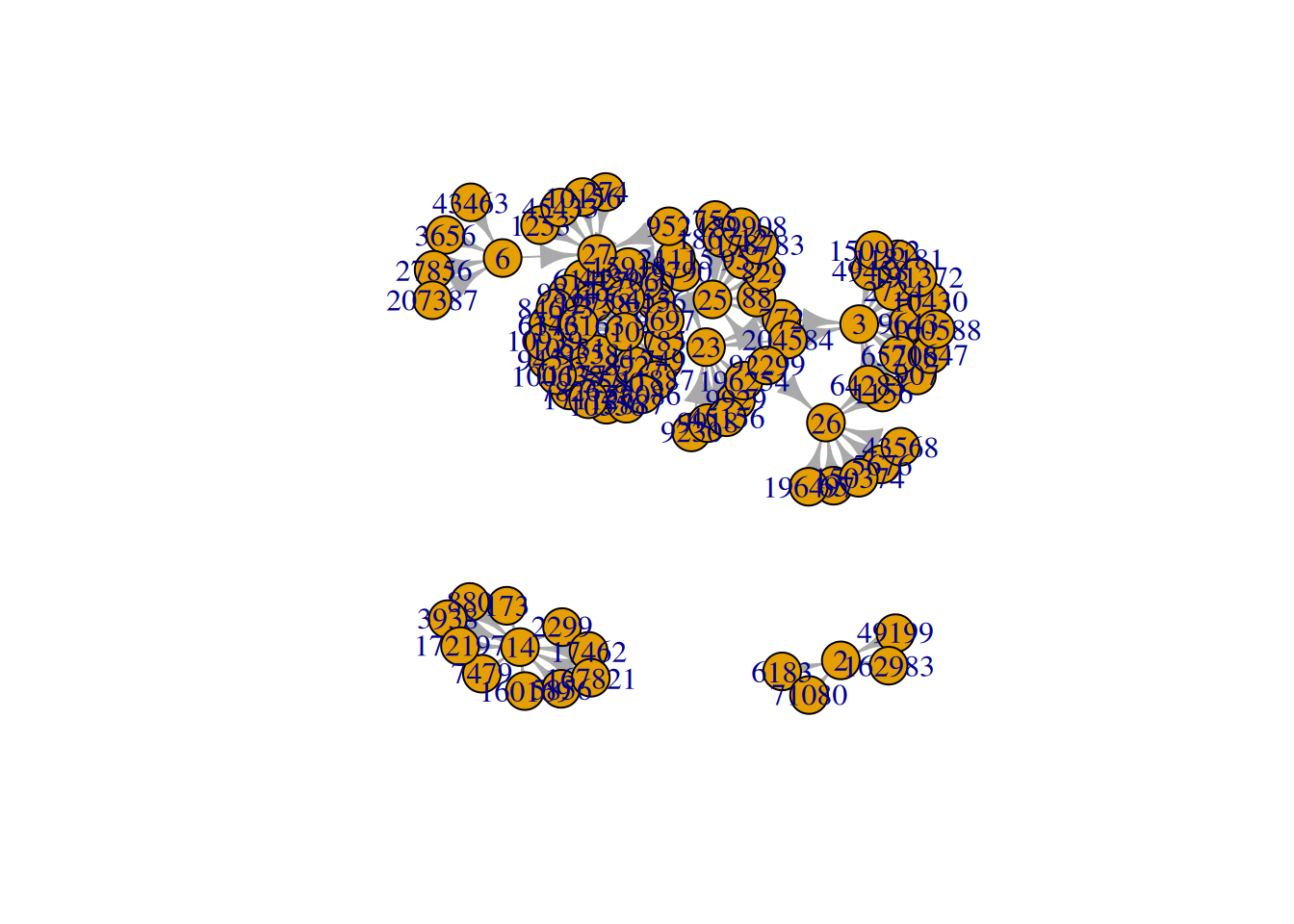
A tk application can launched to show the plot interactively:
plot(g, layout = layout.fruchterman.reingold)Some metrics:
metrics <- data.frame(
deg = degree(g),
bet = betweenness(g),
clo = closeness(g),
eig = evcent(g)$vector,
cor = graph.coreness(g)
)Warning: `evcent()` was deprecated in igraph 2.0.0.
ℹ Please use `eigen_centrality()` instead.Warning: `graph.coreness()` was deprecated in igraph 2.0.0.
ℹ Please use `coreness()` instead.#
head(metrics) deg bet clo eig cor
6183 1 0 1.0000000 5.672735e-17 1
49199 1 0 1.0000000 5.157032e-17 1
71080 1 0 1.0000000 5.672735e-17 1
162983 1 0 1.0000000 4.641329e-17 1
772 3 0 0.3333333 1.040854e-01 2
907 1 0 1.0000000 8.141832e-03 1The metrics above can be interpreted as follows:
degree: local connectivity of each nodebetweenness: how frequently a node lies on shortest pathscloseness: how quickly a node can reach other nodeseig: influence based on connections to influential nodescor: core-periphery position in the network
For better graph readability, filter to the largest connected component or top nodes by degree before plotting labels.
library(ggplot2)
ggplot(
metrics,
aes(x=bet, y=eig,
label=rownames(metrics),
colour=res, size=abs(res))
)+
xlab("Betweenness Centrality")+
ylab("Eigenvector Centrality")+
geom_text()
+
theme(title="Key Actor Analysis")
V(g)$label.cex <- 2.2 * V(g)$degree / max(V(g)$degree)+ .2
V(g)$label.color <- rgb(0, 0, .2, .8)
V(g)$frame.color <- NA
egam <- (log(E(g)$weight)+.4) / max(log(E(g)$weight)+.4)
E(g)$color <- rgb(.5, .5, 0, egam)
E(g)$width <- egam
# plot the graph in layout1
plot(g, layout=layout1)Further information:
http://sna.stanford.edu/lab.php?l=1